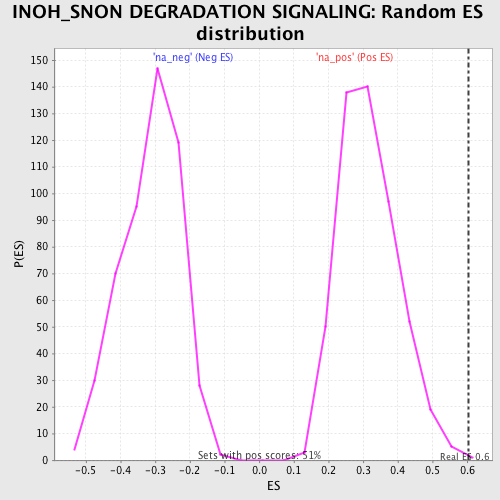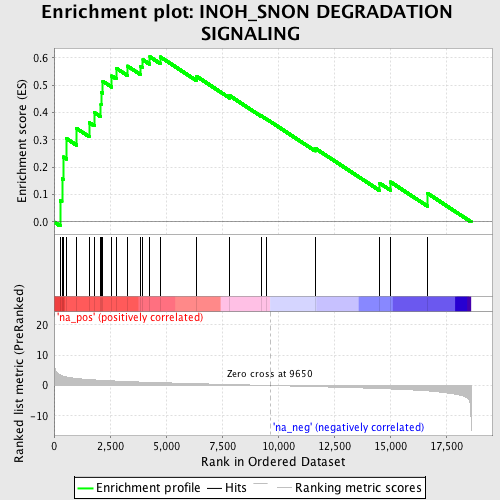
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set04_transDMproB_versus_LMproB |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | INOH_SNON DEGRADATION SIGNALING |
| Enrichment Score (ES) | 0.6034775 |
| Normalized Enrichment Score (NES) | 1.9039736 |
| Nominal p-value | 0.001980198 |
| FDR q-value | 0.033454344 |
| FWER p-Value | 0.385 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | PSMB1 | 271 | 3.438 | 0.0775 | Yes | ||
| 2 | TLE4 | 353 | 3.194 | 0.1586 | Yes | ||
| 3 | PSMB6 | 406 | 3.058 | 0.2377 | Yes | ||
| 4 | PSMB10 | 554 | 2.796 | 0.3046 | Yes | ||
| 5 | PSMB9 | 1005 | 2.291 | 0.3417 | Yes | ||
| 6 | SMAD2 | 1568 | 1.934 | 0.3633 | Yes | ||
| 7 | PSMB2 | 1796 | 1.824 | 0.3999 | Yes | ||
| 8 | PSMB3 | 2090 | 1.688 | 0.4294 | Yes | ||
| 9 | PSMA5 | 2104 | 1.682 | 0.4737 | Yes | ||
| 10 | PSMB8 | 2176 | 1.657 | 0.5142 | Yes | ||
| 11 | PSMA3 | 2575 | 1.524 | 0.5336 | Yes | ||
| 12 | TLE3 | 2771 | 1.457 | 0.5621 | Yes | ||
| 13 | TLE6 | 3270 | 1.298 | 0.5701 | Yes | ||
| 14 | PSMA4 | 3845 | 1.130 | 0.5695 | Yes | ||
| 15 | PSMB5 | 3966 | 1.100 | 0.5925 | Yes | ||
| 16 | PSMA6 | 4276 | 1.032 | 0.6035 | Yes | ||
| 17 | PSMD2 | 4743 | 0.926 | 0.6032 | No | ||
| 18 | PSMA2 | 6367 | 0.596 | 0.5319 | No | ||
| 19 | PSMB4 | 7842 | 0.322 | 0.4612 | No | ||
| 20 | PSMA7 | 9241 | 0.077 | 0.3881 | No | ||
| 21 | PSMB7 | 9486 | 0.035 | 0.3759 | No | ||
| 22 | TLE2 | 11653 | -0.378 | 0.2695 | No | ||
| 23 | PSMA1 | 14534 | -1.013 | 0.1417 | No | ||
| 24 | PSMC1 | 15017 | -1.140 | 0.1463 | No | ||
| 25 | TLE1 | 16668 | -1.763 | 0.1048 | No |